|
R&D Systems
ap2 gamma Ap2 Gamma, supplied by R&D Systems, used in various techniques. Bioz Stars score: 94/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/ap2 gamma/product/R&D Systems Average 94 stars, based on 1 article reviews
ap2 gamma - by Bioz Stars,
2026-04
94/100 stars
|
Buy from Supplier |
|
Sino Biological
human tfap2c cdna  Human Tfap2c Cdna, supplied by Sino Biological, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/human tfap2c cdna/product/Sino Biological Average 90 stars, based on 1 article reviews
human tfap2c cdna - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Proteintech
anti tfap2c  Anti Tfap2c, supplied by Proteintech, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/anti tfap2c/product/Proteintech Average 93 stars, based on 1 article reviews
anti tfap2c - by Bioz Stars,
2026-04
93/100 stars
|
Buy from Supplier |
|
R&D Systems
goat monoclonal tfap2c 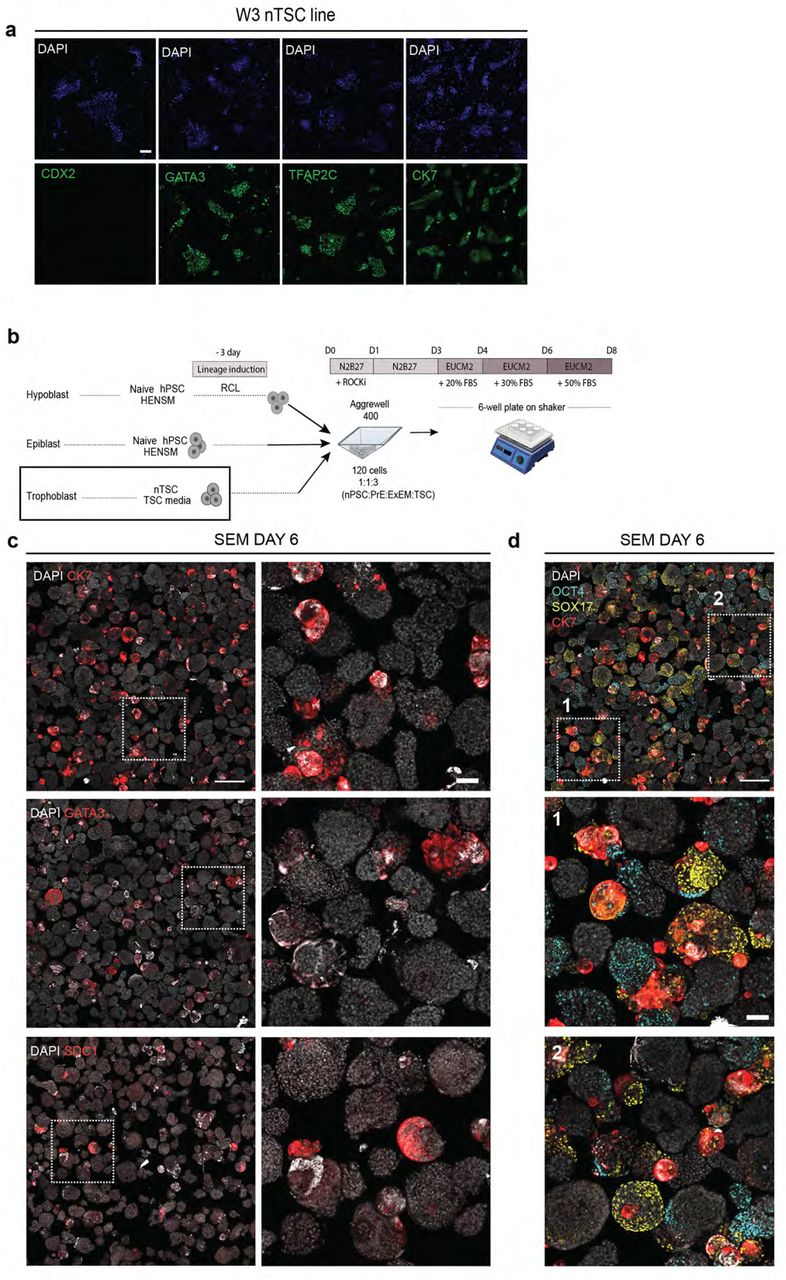 Goat Monoclonal Tfap2c, supplied by R&D Systems, used in various techniques. Bioz Stars score: 94/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/goat monoclonal tfap2c/product/R&D Systems Average 94 stars, based on 1 article reviews
goat monoclonal tfap2c - by Bioz Stars,
2026-04
94/100 stars
|
Buy from Supplier |
|
OriGene
tfap2c sirna  Tfap2c Sirna, supplied by OriGene, used in various techniques. Bioz Stars score: 92/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/tfap2c sirna/product/OriGene Average 92 stars, based on 1 article reviews
tfap2c sirna - by Bioz Stars,
2026-04
92/100 stars
|
Buy from Supplier |
|
OriGene
tfap2c gfp tagged  Tfap2c Gfp Tagged, supplied by OriGene, used in various techniques. Bioz Stars score: 92/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/tfap2c gfp tagged/product/OriGene Average 92 stars, based on 1 article reviews
tfap2c gfp tagged - by Bioz Stars,
2026-04
92/100 stars
|
Buy from Supplier |
|
Bioss
bs 6694r  Bs 6694r, supplied by Bioss, used in various techniques. Bioz Stars score: 91/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/bs 6694r/product/Bioss Average 91 stars, based on 1 article reviews
bs 6694r - by Bioz Stars,
2026-04
91/100 stars
|
Buy from Supplier |
|
Novus Biologicals
anti ap 2  Anti Ap 2, supplied by Novus Biologicals, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/anti ap 2/product/Novus Biologicals Average 90 stars, based on 1 article reviews
anti ap 2 - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Proteintech
anti g2 cop antibody  Anti G2 Cop Antibody, supplied by Proteintech, used in various techniques. Bioz Stars score: 85/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/anti g2 cop antibody/product/Proteintech Average 85 stars, based on 1 article reviews
anti g2 cop antibody - by Bioz Stars,
2026-04
85/100 stars
|
Buy from Supplier |
|
Bio-Techne corporation
ap-2 gamma antibody  Ap 2 Gamma Antibody, supplied by Bio-Techne corporation, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/ap-2 gamma antibody/product/Bio-Techne corporation Average 90 stars, based on 1 article reviews
ap-2 gamma antibody - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
Image Search Results

Journal: Cell stem cell
Article Title: TFAP2C- and p63-Dependent Networks Sequentially Rearrange Chromatin Landscapes to Drive Human Epidermal Lineage Commitment
doi: 10.1016/j.stem.2018.12.012
Figure Lengend Snippet: (A) Schematic representation of the piggyBac TetO-TFAP2C inducible expression system in H9 hESC. TRE, tetracycline-responsive element; rtTA, reverse tetracycline transactivator.
Article Snippet:
Techniques: Expressing

Journal: Cell stem cell
Article Title: TFAP2C- and p63-Dependent Networks Sequentially Rearrange Chromatin Landscapes to Drive Human Epidermal Lineage Commitment
doi: 10.1016/j.stem.2018.12.012
Figure Lengend Snippet: (A) Schematic diagram shows the experimental procedure to study whether TFAP2C-induced surface ectoderm progenitor cells (TetO-TFAP2C-D7) can further differentiate into functional keratinocytes (TetO-TFAP2C-KC) in maturation medium.
Article Snippet:
Techniques: Functional Assay

Journal: Cell stem cell
Article Title: TFAP2C- and p63-Dependent Networks Sequentially Rearrange Chromatin Landscapes to Drive Human Epidermal Lineage Commitment
doi: 10.1016/j.stem.2018.12.012
Figure Lengend Snippet: (A) Schematic illustration of the approach to functionally study p63 via CRISPR/Cas9-mediated gene knockout (KO) during TFAP2C-induced epidermal differentiation.
Article Snippet:
Techniques: CRISPR, Gene Knockout

Journal: Cell stem cell
Article Title: TFAP2C- and p63-Dependent Networks Sequentially Rearrange Chromatin Landscapes to Drive Human Epidermal Lineage Commitment
doi: 10.1016/j.stem.2018.12.012
Figure Lengend Snippet: (A) Gene expression changes of TFAP2C and p63 during TFAP2C-induced epidermal differentiation.
Article Snippet:
Techniques: Expressing

Journal: Cell stem cell
Article Title: TFAP2C- and p63-Dependent Networks Sequentially Rearrange Chromatin Landscapes to Drive Human Epidermal Lineage Commitment
doi: 10.1016/j.stem.2018.12.012
Figure Lengend Snippet: A proposed model depicts the identified chromatin states and feedback regulation between TFAP2C- and p63-centered TF regulatory networks driving the chromatin transition during epidermal lineage commitment.
Article Snippet:
Techniques:

Journal: Cell stem cell
Article Title: TFAP2C- and p63-Dependent Networks Sequentially Rearrange Chromatin Landscapes to Drive Human Epidermal Lineage Commitment
doi: 10.1016/j.stem.2018.12.012
Figure Lengend Snippet: KEY RESOURCES TABLE
Article Snippet:
Techniques: Purification, Recombinant, Knock-Out, Blocking Assay, Plasmid Preparation, Clone Assay, SYBR Green Assay, Magnetic Beads, Staining, Sequencing, CRISPR, Negative Control, shRNA, Expressing, Software

Journal: bioRxiv
Article Title: Transgene-Free Ex Utero Derivation of A Human Post-Implantation Embryo Model Solely from Genetically Unmodified Naïve PSCs
doi: 10.1101/2023.06.14.544922
Figure Lengend Snippet: a, immunofluorescence images validating correct expression of TSC marker genes in colonies of a WIBR3 (W3) naïve hESC derived TSC line (termed nTSC). CK7, TFAP2C, GATA3, CDX2 (all in green); DAPI (blue). Scale bar, 100 µm. b, scheme for aggregation protocol of naïve pluripotent stem cells (nPSCs) in HENSM media, naïve-derived trophoblast stem cells (nTSCs), and nPSCs induced in RCL towards PrE/ExEM for 3 days. c , immunofluorescence images showing rare expression of CK7, GATA3, and SDC1 (red) trophoblast markers in the aggregates. nuclei (DAPI, white); Left column scale bar, 500 µm; Right, zoom into several aggregates with CK7 expression; scale bar, 100 µm. d , immunofluorescence image from (upper left panel in c) , showing Epi (OCT4, cyan) and PrE (SOX17, yellow) with CK7 (red); nuclei (DAPI, white). Zoom ins from the indicated regions are shown. Although some aggregates express lineage markers, they do not organize into embryoid-like structures and are not uniformly surrounded by the trophoblast compartment. Scale bar, 500 µm; bottom, 100 µm.
Article Snippet: The antibodies and dilutions employed for cell immunofluorescence were the following: Rabbit polyclonal anti-Cdx2 (Cell Signaling Cat# 3977) 1:200; Mouse monoclonal anti-Cdx2 (Biogenex Cat# MU392A-UC) 1:200; Rabbit polyclonal anti Gata4 (Abcam Cat# Ab84593) 1:120; Rabbit monoclonal anti-Foxa2 (Abcam Cat# Ab108422) 1:100; Goat polyclonal anti-Sox17 (R&D Cat# AF1924) 1:200; Mouse monoclonal anti-Oct4 (clone C-10) (Santa Cruz Cat# SC-5279) 1:200; Rabbit monoclonal Cdx2 (Abcam Cat# ab76541) 1:200;
Techniques: Immunofluorescence, Expressing, Marker, Derivative Assay

Journal: bioRxiv
Article Title: Transgene-Free Ex Utero Derivation of A Human Post-Implantation Embryo Model Solely from Genetically Unmodified Naïve PSCs
doi: 10.1101/2023.06.14.544922
Figure Lengend Snippet: a, scheme of the donor plasmid vector used for genomic integration of the DOX-inducible iGATA3 overexpression transgene. b , immunofluorescence images of iGATA3 cells, showing uniform GATA3 expression (green) only in response to DOX; nuclei (DAPI, blue). Scale bars, 100 µm. c , brightfield images of WT (top) and iGATA3 (bottom) cells after incubation in BAP(J) media for three days. Scale bar, 200 µm. d , qRT-PCR gene expression (normalized by GAPDH and ACTIN) of the trophoblast markers for WT naïve pluripotent stem cells (nPSCs) in BAP(J) media (purple) and iGATA3 nPSC cells induced by DOX in BAP(J) media (yellow), versus nPSCs maintained in HENSM media and used as a reference control (set as 1) (white). e , immunofluorescence images showing different patterns of expression of trophoblast marker genes in the wild type (WT) nPSCs incubated in BAP(J) media for three days (top) versus iGATA3 cells, induced by DOX in BAP(J) media (bottom). GATA3 (magenta), TFAP2C (magenta), GATA2, CDX2, SDC1, and HCGB (all in green), nuclei (DAPI, blue). Scale bar, 100 µm. f , immunofluorescence images showing uniform expression of CK7 in colonies of WT nPSCs incubated in BAP(J) media for three days (top), whereas colonies of iGATA3 cells induced by DOX in BAP(J) media express CK7 heterogeneously, with most of them being negative for CK7. Scale bar, 100 µm. g , from left to right, FACS plots of ENPEP versus TACSTD2 for trophoblast (Tb) priming using iGATA3 induction in different media (AP(J) with and without DOX, BAP(J) with and without DOX), and using WT nPSC priming to trophectoderm in AP(J) and BAP(J) regimens. Percentage of double positive population is indicated. This result shows that transient expression of the GATA3 transgene is not required for Tb differentiation and that it can be achieved with BAP(J) media using WT nPSCs as starting cells.
Article Snippet: The antibodies and dilutions employed for cell immunofluorescence were the following: Rabbit polyclonal anti-Cdx2 (Cell Signaling Cat# 3977) 1:200; Mouse monoclonal anti-Cdx2 (Biogenex Cat# MU392A-UC) 1:200; Rabbit polyclonal anti Gata4 (Abcam Cat# Ab84593) 1:120; Rabbit monoclonal anti-Foxa2 (Abcam Cat# Ab108422) 1:100; Goat polyclonal anti-Sox17 (R&D Cat# AF1924) 1:200; Mouse monoclonal anti-Oct4 (clone C-10) (Santa Cruz Cat# SC-5279) 1:200; Rabbit monoclonal Cdx2 (Abcam Cat# ab76541) 1:200;
Techniques: Plasmid Preparation, Over Expression, Immunofluorescence, Expressing, Incubation, Quantitative RT-PCR, Marker

Journal: PLoS ONE
Article Title: Transcription Factor TFAP2C Regulates Major Programs Required for Murine Fetal Germ Cell Maintenance and Haploinsufficiency Predisposes to Teratomas in Male Mice
doi: 10.1371/journal.pone.0071113
Figure Lengend Snippet: Affymetrix microarray gene expression analysis performed with RNA extracted from #1- ctrl ; #1- Tfap2c −/− and #2- ctrl ; #2- Tfap2c −/− ESCs and PGCLCs. (A) Hierarchical clustering. The red bars cluster the ESCs, green bars the Tfap2c −/− PGCLCs and blue bars ctrl PGCLCs, respectively. The shorter the horizontal bar that connects two branches the closer are the populations. (B) Heat map was performed with the probes whose range or variation across all samples was at least 3. Color bar on top codifies the gene expression in log2 scale. Red and blue indicate higher and lower relative expression. (C-D) Pairwise scatter plot of global gene expression in ctrl versus Tfap2c −/− PGCLCs (C) and ESCs (D). Black lines indicate 1.5 fold-change in log2 scale of gene expression levels between paired PGCLCs and ESCs. Color bars on the side display the scattering density with light blue indicating lower and blue higher scatter density. Genes upregulated in Tfap2c −/− samples are shown as red dots; genes downregulated are shown as green dots. R 2 = Fisher’s correlation coefficient. (E) Venn diagram; in PGCLCs 455 genes are deregulated; in ESCs 26 genes are deregulated. The intersection part show the commonly deregulated genes (n = 13) by TFAP2C in PGCLCs and ESCs (Fold-change >1.5 in log2 scale).
Article Snippet: Scrambled and
Techniques: Microarray, Expressing

Journal: PLoS ONE
Article Title: Transcription Factor TFAP2C Regulates Major Programs Required for Murine Fetal Germ Cell Maintenance and Haploinsufficiency Predisposes to Teratomas in Male Mice
doi: 10.1371/journal.pone.0071113
Figure Lengend Snippet: (A) Quantitative RT-PCR of a subset of markers with RNA isolated from ctrl and Tfap2c −/− PGCLCs. Expression levels were normalized to βActin and expression level of ctrl PGCLCs were set to 1. qRT-PCR was performed in biological triplicates. Error bars indicate standard deviation. (B) Quantitative RT-PCR of a subset of markers with RNA isolated from TCam-2 cells after siRNA mediated knockdown of TFAP2C. Expression levels were normalized to GAPDH and expression levels of scrambled-siRNA-transfection were set to 1. qRT-PCR was performed in biological triplicate. Error bars indicate standard deviation. (C) ChIP/qPCR analysis for Tfap2c was performed with four biological replicates of PGCLCs. The qPCR results were calculated with the percentage input method and ChIP analyses with IgG antibody served as control and were set to 1. Error bars indicate standard deviation. ChIP analysis demonstrates increased binding of TFAP2C at indicated loci. (A) – (C) Marker genes were grouped in categories: somatic differentiation (category I), germ cell maintenance and maturation (category II), cell cycle regulation (category III), pluripotency (category IV) and epigenetic modification (category V).
Article Snippet: Scrambled and
Techniques: Quantitative RT-PCR, Isolation, Expressing, Standard Deviation, Transfection, Binding Assay, Marker, Modification

Journal: PLoS ONE
Article Title: Transcription Factor TFAP2C Regulates Major Programs Required for Murine Fetal Germ Cell Maintenance and Haploinsufficiency Predisposes to Teratomas in Male Mice
doi: 10.1371/journal.pone.0071113
Figure Lengend Snippet: (A) Quantitative RT-PCR with RNA isolated from E12.5 genital ridges of wt and Tfap2c +/− embryos was performed. Expression levels of Tfap2c and p21 were normalized to Gapdh . qRT-PCR was performed in biological duplicates. Error bars indicate standard deviation. (B) Percentage of total tumor incidence in Tfap2c heterozygous 129S2/Sv male mice. The seventh generation in 129S2/Sv shows 82% (n = 51) testicular tumors. Red bar: bilateral cases (35%), blue bar: unilateral tumors (47%). (C) Gross pathology of testicular teratoma in Tfap2c +/− male mice. (D–G) HE-staining of testicular teratomas of 3–6 month old mice. Tumors show immature glia (D), mature cartilage, muscle (E), respiratory epithelium (F) and squamous epithelium (G). Scale bars: 200 µm.
Article Snippet: Scrambled and
Techniques: Quantitative RT-PCR, Isolation, Expressing, Standard Deviation, Staining

Journal: PLoS ONE
Article Title: Transcription Factor TFAP2C Regulates Major Programs Required for Murine Fetal Germ Cell Maintenance and Haploinsufficiency Predisposes to Teratomas in Male Mice
doi: 10.1371/journal.pone.0071113
Figure Lengend Snippet: (A–C) IHC staining of E16.5 embryonal testis. (A) SSEA1, (B) OCT3/4 and (C) TFAP2C. Scale bars: 50 µm. (B) Black arrow show cells with large, highly condensated nuclei as well as clearly visible nucleoli. (D) RT-PCR from RNA of microdissected SSEA-1 positive foci of E16.5 testes detecting Nanog , Sox2 and c-Kit . Comparison of microdissected SSEA-1 positive tissue (T) and SSEA-1 negative tissue (wt). Gapdh served as control.
Article Snippet: Scrambled and
Techniques: Immunohistochemistry, Reverse Transcription Polymerase Chain Reaction

Journal: PLoS ONE
Article Title: Transcription Factor TFAP2C Regulates Major Programs Required for Murine Fetal Germ Cell Maintenance and Haploinsufficiency Predisposes to Teratomas in Male Mice
doi: 10.1371/journal.pone.0071113
Figure Lengend Snippet: Black arrows indicate pathways transactivated and induced by TFAP2C and black lines with terminal bars indicate pathways repressed by TFAP2C during development of primordial germ cells. Genes listed in respective pathways indicate direct regulation as demonstrated by ChIP analyses.
Article Snippet: Scrambled and
Techniques:

Journal: Frontiers in Cell and Developmental Biology
Article Title: ESRRB Facilitates the Conversion of Trophoblast-Like Stem Cells From Induced Pluripotent Stem Cells by Directly Regulating CDX2
doi: 10.3389/fcell.2021.712224
Figure Lengend Snippet: Antibodies used in this experiment.
Article Snippet: CDX2 ,
Techniques: